Prognostic Plant Traits
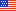
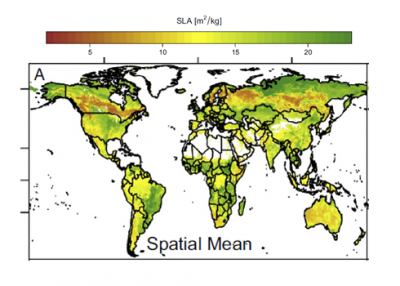
Variation in plant traits at local scales can be very large, of similar magnitude to the global range of values used to characterize mean traits for multiple plant functional types. We generated global maps of plant trait distributions for use in Earth system models, representing trait means and trait variation in each global grid cell. We use environmental variables to estimate trait distributions so that trait variation under future conditions can be estimated as well.
We use the largest available global plant traits database and global environmental datasets as observational inputs to a Bayesian prediction framework. We estimated trait distributions (mean and standard deviation) for three leaf traits known to play a critical role in predictions of photosynthesis and respiration: specific leaf area, leaf nitrogen concentration, and leaf phosphorus concentration. We made global trait distribution estimates both with and without constraints from global maps of plant functional type distribution.
Currently, Earth system models (ESMs) represent variation in plant life through the presence of a small set of plant functional types (PFTs), each of which accounts for hundreds or thousands of species across thousands of vegetated grid cells on land. By expanding plant traits from a single mean value per PFT to a full distribution per PFT that varies among grid cells, the trait variation present in nature is restored and may be propagated to estimates of ecosystem processes. Indeed, critical ecosystem processes tend to depend on the full trait distribution, which therefore needs to be represented accurately. These maps reintroduce substantial local variation and will allow for a more accurate representation of the land surface in ESMs.
The global land surface is perhaps the most heterogeneous component of the Earth system. Reducing vegetation to a collection of plant functional types (PFTs) with fixed trait values has been the preferred
method to constrain this heterogeneity and group similar biochemical and biophysical properties; however, this has been at the expense of functional diversity. This analysis quantifies
the substantial magnitude of this ignored trait variation. The approach and methods presented here retain the simplicity of the PFT representation, but capture a wider range of functional
diversity.
This research was supported as part of the Energy Exascale Earth System Model (E3SM) project, funded by the US Department of Energy, Office of Science, Office of Biological and Environmental Research (Grant DE-SC0012677).